VIVO Documentation
Old Release
This documentation relates to an old version of VIVO, version 1.10.x. Looking for another version? See all documentation.
Overview
VIVO provides the capability to add ontologies and use them to describe your data and for query. You can add your own, or pre-existing ontologies. In each case, the ontology to be added will appear in the VIVO Ontology List along with ontologies provided with VIVO.
If your ontology assertions contain an owl:Ontology assertion giving the URI of your ontology, your ontology will appear in Site Admin → Ontology List. Otherwise, your ontology assertions will be added to be VIVO and will be fully functional as classes, object properties, and data properties, but will not be associated with a named ontology.
Ontologies added to VIVO should use the .owl file type, and should be written in OWL DL. Ontologies written in RDF or OWL FULL are not supported by VIVO.
Using the Web Interface
Create an Ontology from the Web Interface
To create an ontology from the web interface, follow the steps in Create, Assign, and Use an Institutional Internal Class. These instructions show how to create an ontology using the web interface. Once your ontology is created, you can add an Institutional Class as described, additional classes, data properties and Object properties. You may create as many additional ontologies as you need.
Load an Existing Ontology as Data
You can add an ontology from a file into VIVO using the Add/Remove RDF Data feature shown on the Site Admin page. This method is simple, but not recommended for production sites. Your ontology will be loaded into the VIVO main data graph and could be very difficult to remove. Much preferred for production sites is the following method, Loading to a Named Graph.
Load an Existing Ontology to a Named Graph
Use Site Admin → Ingest Tools → Manage Jena Models → Create Jena Model
Give your named graph (Jena calls them models) a name such as myOntology.owl
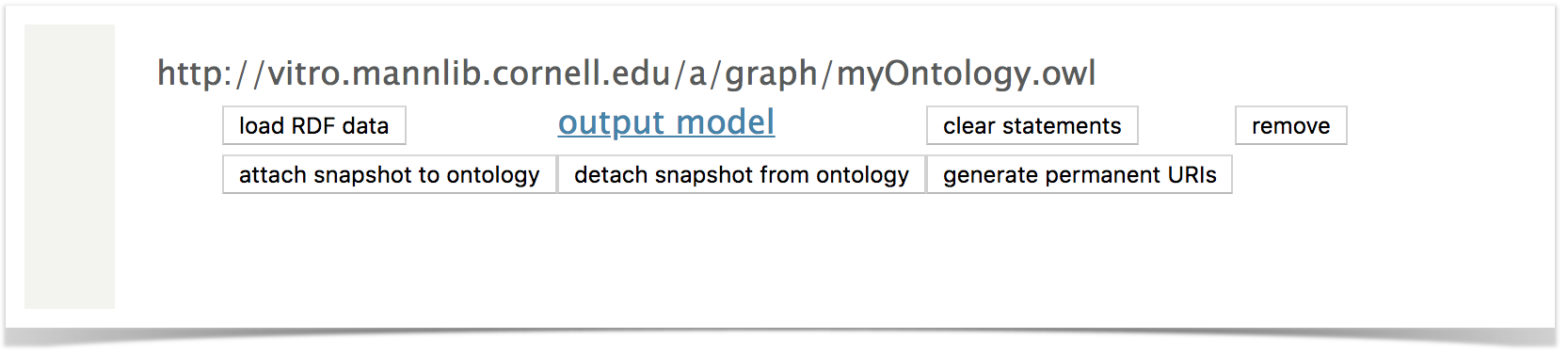
You will see your ontology represented as an empty named graph in the list of Graphs.
Click Load RDF data, and upload your ontology file to your named graph.
In this way, you keep your ontology assertions separate from other VIVO assertions. You can drop the graph, and/or reload it as needed.
Loading from Filegraphs
There is another mechanism for incorporating ontologies into VIVO. This involves "filegraphs," and is how the VIVO ontology is included with the software. Filegraphs are RDF documents stored in the VIVO home directory. Each filegraph corresponds to a single named graph in the triple store. Every time Tomcat starts, VIVO checks each of these graphs to ensure that its contents exactly match the triples found in the corresponding file. If the file has changed, VIVO makes the necessary modifications to the corresponding named graph in the triple store. If a filegraph is removed from its directory, its graph in the triple store will be deleted the next time Tomcat starts.
Example
vivo.home/
rdf/
tbox/
filegraph/
vivo.owl
ontologies.owl
...
myOntology.owl
...
Adding myOntology.owl to the directory as shown above will automatically create the corresponding named graph in the triple store after Tomcat is restarted:
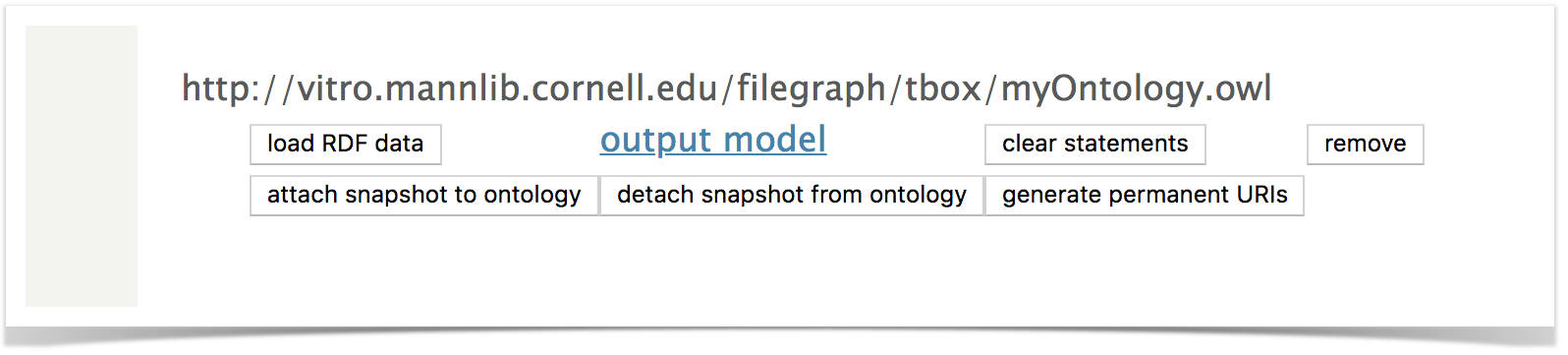
After restart, go to Site Admin → Ingest Data → Manage Jena Models. Review the named graphs. You should see:
Removing the myOntology.owl file from the filegraph directory and restarting Tomcat will automatically remove the named graph. This will remove all ontology assertions made in myOntology.owl. It will not remove any abox assertions using myOntology.owl from the content triple store.
Namespace Prefixes
owl:Ontology statements result in the ontology being added to VIVO's Ontology List. You might see an entry such as:
Indicating that the ontology you have added does not have a label or prefix.
Specify a label and a prefix for your ontology as shown below:
<owl:Ontology rdf:about="http://any.ontology.org/core/">
<rdfs:label xml:lang="en-US">My Ontology</rdfs:label>
<vitro:ontologyPrefixAnnot xml:lang="en-US">myo</vitro:ontologyPrefixAnnot>
</owl:Ontology>
Restarting Tomcat to reload the filegraph will display your prefix and label.